WEPP: Wastewater based Epidimic using phylogenetic placement
If the last few years have taught us anything, it’s that wastewater has a story to tell. During COVID-19, scientists realized that the stuff flowing through our pipes carries fragments of viral genomes shed by everyone in the community. With the right tools, that messy mix can give us an early warning of outbreaks — sometimes weeks before clinical testing catches up. That’s the idea behind wastewater-based epidemiology (WBE), and it’s been adopted everywhere from university dorms to entire cities.
But here’s the catch: most existing tools only get us so far. They can usually tell us which lineages (like BA.5, XBB, etc.) are in a sample and roughly how abundant they are. That’s useful, but it’s a bit like knowing which family someone belongs to without knowing which sibling they actually are. When you’re trying to track outbreaks, introductions, or novel variants, lineage-level resolution just isn’t enough.
In collaboration with the Turakhia Lab at UC San Diego, we developed WEPP: Wastewater Epidemiology using Phylogenetic Placements
What is WEPP?
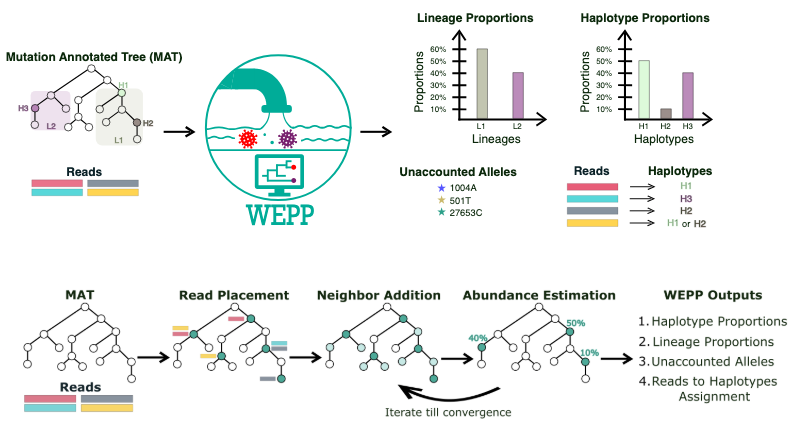
WEPP stands for Wastewater Epidemiology using Phylogenetic Placements. Instead of stopping at lineages, WEPP zooms in on haplotypes — the actual sequence variants floating around in a community. It works by placing wastewater sequencing reads onto a mutation-annotated phylogenetic tree (MAT) — essentially a giant family tree of all the clinical sequences that have ever been collected (plus inferred ancestors).
From there, WEPP figures out which haplotypes are most likely present in the wastewater, estimates their abundances, and even highlights mutations it can’t explain (these “unaccounted alleles” often hint at cryptic or novel variants).
And the best part? WEPP isn’t just for SARS-CoV-2. It’s designed to be pathogen-agnostic, and the team has already tested it on things like RSV and Mpox.
WEPP stands for Wastewater Epidemiology using Phylogenetic Placements. Instead of stopping at lineages, WEPP zooms in on haplotypes — the actual sequence variants floating around in a community. It works by placing wastewater sequencing reads onto a mutation-annotated phylogenetic tree (MAT) — essentially a giant family tree of all the clinical sequences that have ever been collected (plus inferred ancestors).
From there, WEPP figures out which haplotypes are most likely present in the wastewater, estimates their abundances, and even highlights mutations it can’t explain (these “unaccounted alleles” often hint at cryptic or novel variants).
And the best part? WEPP isn’t just for SARS-CoV-2. It’s designed to be pathogen-agnostic, and the team has already tested it on things like RSV and Mpox.

Why does this matter?
By moving beyond lineage-level summaries, WEPP makes it possible to do things that older wastewater tools couldn’t. It can distinguish fine-grained clusters within a single lineage, helping reveal separate introductions or localized outbreaks that would otherwise be hidden. It can also flag when a new variant cluster shows up in a region and even point to its likely source, providing an early signal for public health action. On top of that, WEPP systematically surfaces unexplained mutations—so-called unaccounted alleles—that often mark the presence of cryptic or emerging variants. And because it’s not limited to SARS-CoV-2, WEPP extends these capabilities to pathogens like RSV, Mpox, and influenza, where clinical sequencing data are often scarce.
A dashboard for exploration
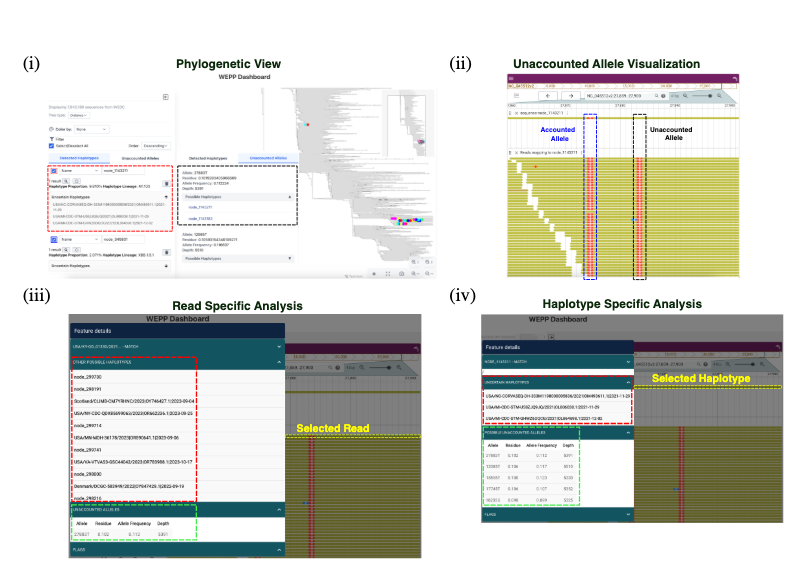
Another neat part of WEPP is its interactive dashboard. It’s not just a black-box pipeline — you can actually visualize:- Haplotypes detected in your sample on the global phylogeny
- Their estimated abundances and lineage assignments
- Read-to-haplotype mappings
- The mysterious unaccounted alleles
It's built on top of Taxonium, but built specifically for wastewater data. The public health teams don’t just get summary stats — they can actually dig into the details and interpret what’s going on.

Wastewater is one of the most promising tools we have for post-pandemic pathogen surveillance, especially as clinical sequencing efforts taper off. With WEPP, we now have a way to use that signal at near-haplotype resolution, not just lineages. That opens up possibilities for faster detection, finer-scale tracking, and broader pathogen coverage.